class: center, middle, inverse, title-slide # Single Cell Analysis and Visualization ## Lab Meeting: 2019-04-12 ### Kevin Stachelek ### 2019/04/08 (updated: 2019-04-12) --- ### What are our resources for computing in the lab? We have two computers with the power to run pipelines and host apps -- .pull-left[ Thor (ip address 10.134.4.240) <img src="img/tower.png" width="200" height="200" /> ] -- .pull-right[ SRT server (ip address 10.134.4.105) <img src="img/server.png" width="200" height="200" /> ] --- ### We also have one back up server located on the USC campus USC server (ip address 10.112.120.117) <img src="img/server.png" width="200" height="200" /> --- ### How can we use these resources? + Use [rstudio](10.134.4.105:8787) on one of your servers via your web browser -- + Connect in a terminal -- + Use a graphical user interface (gui) that manages this connection for you: 1. Cyberduck (Mac) 2. filezilla (Mac/Windows) 3. PuTTY (Windows) -- + Use 'Shiny' Web Apps -- + Use jupyter notebooks (python) --- ### Projects - Transcriptional Response to Rb Loss in the Retinoblastoma Cell of Origin Single Cell RNA Sequencing (scRNAseq) - Changes associated with retinal cone precursor development Single Cell RNA Sequencing (scRNAseq) --- ### What are the transcriptomic changes involved in Retinoblastoma Tumorigenesis? <img src="img/rb_pathway_diagram.png" width="900px" /> --- ### Study Design <img src="img/single_cell_exp_scheme.png" width="800" height="400" /> --- ### Analysis Pipeline <img src="img/single_cell_pipeline_flowchart.png" width="300" height="450" /> --- ### Plot hierarchical clustering <img src="img/Capture.PNG" width="900" style="display: block; margin: auto;" /> --- ### Calculate Principle Components <style> #wrap { width: 900px; height: 1000px; padding: 0; overflow: hidden; } #frame { width: 800px; height: 700px; border: 0px solid black; } #frame { -ms-zoom: 0.75; -moz-transform: scale(0.75); -moz-transform-origin: 0 0; -o-transform: scale(0.75); -o-transform-origin: 0 0; -webkit-transform: scale(0.75); -webkit-transform-origin: 0 0; } </style> <div id="wrap"> <iframe id="frame" frameborder="0" scrolling="no" src="img/Plot_1.html"></iframe> </div> --- ### Plot Trajectory <iframe width="900" height="750" frameborder="0" scrolling="no" src="img/shRB_cluster_colors2.html"></iframe> --- ### Calculate Pseudotime <iframe width="900" height="750" frameborder="0" scrolling="no" src="img/PT_shRB_shCtrl2.html"></iframe> --- ### Find Genes Correlated with Pseudotime <img src="img/spearman_corr_1.PNG" width="900px" /><img src="img/spearman_corr_2.PNG" width="900px" /> --- ### Why did this not work well? -- + Normalization -- + Batch Effects -- + Incomplete Dimensional Reduction -- + Naive Clustering Approach --- ### The Next Generation: Seurat #### What is Seurat? Seurat is an R package designed for QC, analysis, and exploration of single-cell RNA-seq data. Written in R so fits easily into existing analysis --- ### Dimensional Reduction by PCA <img src="img/pca.gif" height="400" /> --- ### Graph Construction and Clustering <img src="img/phenograph.jpg" height="400" /> --- ### Further Dimensional Reduction #### Several Techniques: most common is tSNE or UMAP <img src="img/tsne_v_umap.gif" width="960" height="480" /> --- ### Seurat Advantages: 1. Batch Correction aka 'integration'. 2. Label Transfer across experiments 3. Normalization --- ### Batch Correction aka integration. Seurat v3 implements methods to identify ‘anchors’ across diverse single-cell data types to construct harmonized references, or to transfer information across experiments. Stuart, Butler, Hoffman, Hafemeister, Papalexi, Mauck, Stoeckius, Smibert, and Satija (2018) <img src="img/stuart_integration_diagram.png" width="900" /> --- ### Label Transfer across experiments We can use the same batch correction technique to predict the cluster that a cell from a 'query' dataset would fall into in a reference dataset. Useful for comparison to published studies of retinal transcriptomes #### Mouse Macosko, Basu, Satija, Nemesh, Shekhar, Goldman, Tirosh, Bialas, Kamitaki, Martersteck, Trombetta, Weitz, Sanes, Shalek, Regev, and McCarroll (2015) #### Mouse/Macaque Peng, Shekhar, Yan, Herrmann, Sappington, Bryman, van Zyl, Do, Regev, and Sanes (2019) #### Human Bulk Eldred, Hadyniak, Hussey, Brenerman, Zhang, Chamling, Sluch, Welsbie, Hattar, Taylor, Wahlin, Zack, and Johnston (2018); Mellough, Bauer, Collin, Dorgau, Zerti, Dolan, Jones, Izuogu, Yu, Hallam, Steyn, White, Steel, Santibanez-Koref, Elliott, Jackson, Lindsay, Grellscheid, and Lako (2019) --- ### Normalization Seurat v3 includes sctransform, a new modeling approach for the normalization of single-cell data. Compared to standard log-normalization, sctransform effectively removes technically-driven variation while preserving biological heterogeneity. Hafemeister and Satija (2019) --- ### Other Methods #### [Scanpy](https://scanpy.readthedocs.io/en/latest/) Strengths and Weaknesses 1. Speed 2. Pseudotime Integration - PAGA 3. Makes several machine learning approaches easier to use + Denoising Auto Encoder Eraslan, Simon, Mircea, Mueller, and Theis (2019) + Integrating Datasets (Batch Correction) using Machine Learning Lotfollahi, Wolf, and Theis (2018) + Transfer Learning Lotfollahi, Wolf, and Theis (2018) --- #### [bigScale 2](https://github.com/iaconogi/bigSCale2) * disclaimer: I haven't tried this sofware; and haven't yet gotten a sense of popularity + Sensitive and accurate marker detection and classification. No method is used to reduce dimensions, all information is retained. + Infer gene regulatory networks for any single cell dataset. + Compress large datasets of any size into a smaller datasets of higher quality, without loss of information. + Reduce a dataset of many cells to one with fewers cells of increased quality --- ### So many tools! <blockquote class="bluesky-tweet" data-lang="en"><p lang="en" dir="ltr">Too many awesome new datasets, biological findings, ML methods every second of every day. And they are spawning too many new ideas. I luv science & I'm going crazy (in a good way) that I can't keep up with all the coolness and just have one brain and two hands and no time. Fk!!!!</p>— ANSHUL KUNDAJE (@anshulkundaje) <a href="https://bluesky.com/anshulkundaje/status/1116398619072942080?ref_src=twsrc%5Etfw">April 11, 2019</a></blockquote> <script async src="https://platform.bluesky.com/widgets.js" charset="utf-8"></script> ### What's the most efficient way to pursue our (biological) questions? #### We need to manage our data well and ask the right questions --- ### Step 1: (General) Data Science for all .pull-left[ <span style="font-size: 200%">[R for Data Science](https://r4ds.had.co.nz/)</span> <span style="font-size: 200%">[rstudio cheat sheets](https://www.rstudio.com/resources/cheatsheets/)</span> ] .pull-right[ <!-- --> ] --- # Data Science Workflow 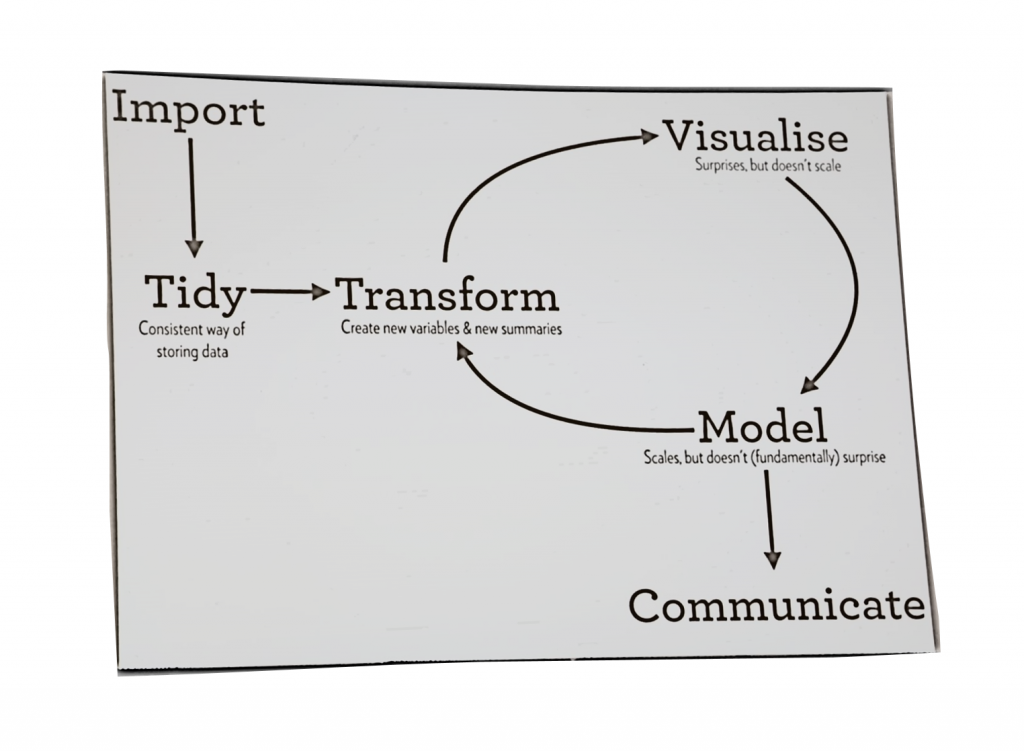<!-- --> --- ### Step 2: Graphical User Interfaces(GUIs) for some #### Shiny & Plotly! --- ### What is Shiny? Shiny is an R package that makes it easy to build interactive web apps straight from R. [Our Apps on Thor (10.134.4.240)](10.134.4.105:3838) [Our Apps on SRT server (10.134.4.105)](10.134.4.105:3838) --- There are more possiblities! Let's think about how graphical tools can help us make progress more quickly [interactive genome browser using Shiny](https://shiny.rstudio.com/gallery/genome-browser.html) --- ### References Eldred, K. C, S. E. Hadyniak, K. A. Hussey, et al. (2018). "Thyroid hormone signaling specifies cone subtypes in human retinal organoids". En. In: _Science_ 362.6411, p. eaau6348. ISSN: 0036-8075, 1095-9203. DOI: [10.1126/science.aau6348](https://doi.org/10.1126%2Fscience.aau6348). URL: [http://www.sciencemag.org/lookup/doi/10.1126/science.aau6348](http://www.sciencemag.org/lookup/doi/10.1126/science.aau6348) (visited on Mar. 26, 2019). Eraslan, G, L. M. Simon, M. Mircea, et al. (2019). "Single-cell RNA-seq denoising using a deep count autoencoder". En. In: _Nature Communications_ 10.1, p. 390. ISSN: 2041-1723. DOI: [10.1038/s41467-018-07931-2](https://doi.org/10.1038%2Fs41467-018-07931-2). URL: [https://www.nature.com/articles/s41467-018-07931-2](https://www.nature.com/articles/s41467-018-07931-2) (visited on Apr. 11, 2019). Hafemeister, C. and R. Satija (2019). "Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression". En. In: _bioRxiv_, p. 576827. DOI: [10.1101/576827](https://doi.org/10.1101%2F576827). URL: [https://www.biorxiv.org/content/10.1101/576827v2](https://www.biorxiv.org/content/10.1101/576827v2) (visited on Apr. 11, 2019). Lotfollahi, M, F. A. Wolf, and F. J. Theis (2018). "Generative modeling and latent space arithmetics predict single-cell perturbation response across cell types, studies and species". En. In: _bioRxiv_, p. 478503. DOI: [10.1101/478503](https://doi.org/10.1101%2F478503). URL: [https://www.biorxiv.org/content/10.1101/478503v2](https://www.biorxiv.org/content/10.1101/478503v2) (visited on Apr. 11, 2019). --- ### References cont. Macosko, E, A. Basu, R. Satija, et al. (2015). "Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets". In: _Cell_ 161.5, pp. 1202-1214. ISSN: 0092-8674. DOI: [10.1016/j.cell.2015.05.002](https://doi.org/10.1016%2Fj.cell.2015.05.002). URL: [http://www.sciencedirect.com/science/article/pii/S0092867415005498](http://www.sciencedirect.com/science/article/pii/S0092867415005498) (visited on Apr. 11, 2019). Mellough, C. B, R. Bauer, J. Collin, et al. (2019). "An integrated transcriptional analysis of the developing human retina". En. In: _Development_ 146.2, p. dev169474. ISSN: 0950-1991, 1477-9129. DOI: [10.1242/dev.169474](https://doi.org/10.1242%2Fdev.169474). URL: [http://dev.biologists.org/lookup/doi/10.1242/dev.169474](http://dev.biologists.org/lookup/doi/10.1242/dev.169474) (visited on Feb. 05, 2019). Peng, Y, K. Shekhar, W. Yan, et al. (2019). "Molecular Classification and Comparative Taxonomics of Foveal and Peripheral Cells in Primate Retina". En. In: _Cell_. ISSN: 00928674. DOI: [10.1016/j.cell.2019.01.004](https://doi.org/10.1016%2Fj.cell.2019.01.004). URL: [https://linkinghub.elsevier.com/retrieve/pii/S0092867419300376](https://linkinghub.elsevier.com/retrieve/pii/S0092867419300376) (visited on Feb. 03, 2019). Stuart, T., A. Butler, P. Hoffman, et al. (2018). _Comprehensive integration of single cell data_. En. preprint. Genomics. DOI: [10.1101/460147](https://doi.org/10.1101%2F460147). URL: [http://biorxiv.org/lookup/doi/10.1101/460147](http://biorxiv.org/lookup/doi/10.1101/460147) (visited on Apr. 11, 2019). --- ### Thanks!